Documentation
At Douglas Connect we focus on improving data management and data access solutions for scientific, industry and regulatory use. A very important effort along these lines was the development of REST style APIs to enable real time access to three of the most widely used toxicological data sources: The EPA's in vitro ToxCast/Tox21 database, the EPA's in vivo ToxRefDB database and the NIBIOHN's toxicogenomics Open TG-Gates database.
The data can be accessed through using the APIs or the user friendly web interface.
Web based Data Explorer
To enable easy browsing of the data served by the APIs, we are currently developing a web based data explorer tool that uses the APIs to show a nice interface to the data, complete with rich filtering and search. With every user operation, the URL is updated that can be used to query for exactly the currently visible data set directly from our APIs.
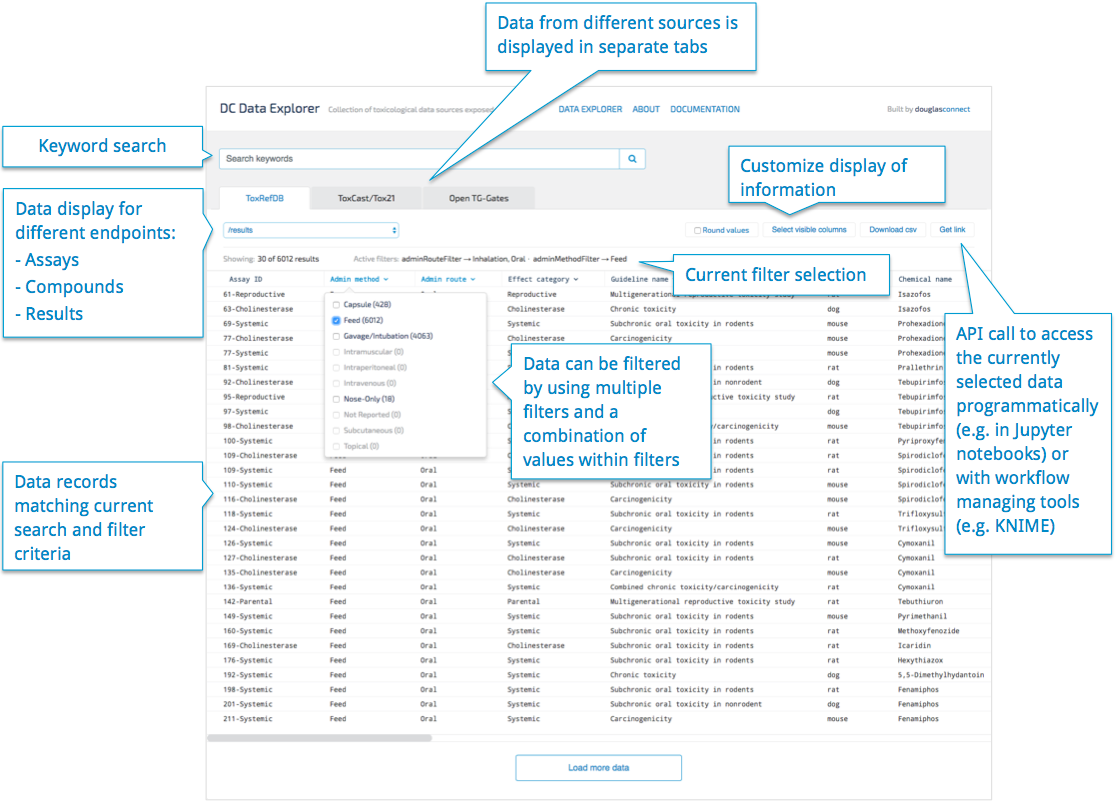
The APIs
By exposing these databases as simple APIs using the widely understood JSON serialisation format, we made it much easier to access their data using a wide variety of tools, from R to Python to KNIME to Javascript running in the browser.
OpenAPI/Swagger definitions
OpenAPI/Swagger definitions are used to document the APIs and enable automated discovery services etc. in the future. The ToxCast/Tox21 and ToxRefDB APIs are currently in public beta with the Open-TGGates expected to follow soon. You can find their documentation here:
GitHub
The ToxCast/Tox21 API is already open source and available at GitHub (including the code for both the API itself and the importer that parses the official upstream data release). The APIs for ToxRefDB and Open TG-Gates will be open sourced in the near future.
The data sources
The EPA's in vitro ToxCast/Tox21 database:
https://www.epa.gov/chemical-research/toxicity-forecaster-toxcasttm-data
The EPA's in vivo ToxRefDB database:
https://www.epa.gov/chemical-research/toxcast-toxrefdb-data-release
The NIBIOHN's toxicogenomics Open TG-Gates database:
http://toxico.nibiohn.go.jp/english/